
how are the physical properties of the cell controlled?
Tens of thousands of biochemical reactions occur simultaneously in the cell. Small molecules are channeled through metabolic pathways at blistering speed. Giant complexes assemble to orchestrate transcription and translation. ATP fuels the active transport of organelles along microtubules, and actin networks drive membrane remodeling and agitate the cytoplasm. All of this occurs within a crowded cell interior that approaches the physical limits where jamming will occur. This extreme physical environment is both essential for life, and a potential liability. If cells become too dilute, they senesce and die. On the other hand, increased crowding eventually stalls growth. We found that the central growth regulator, mTORC1, is a crucial regulator of crowding in both the cytoplasm and nucleus. We also found that mechanical compression affects crowding and phase separation. We are investigating how perturbations to the physical properties of the cell interior through mechanical forces and the regulation of growth pathways impact cell and developmental biology.
The effects of pressure on cancer
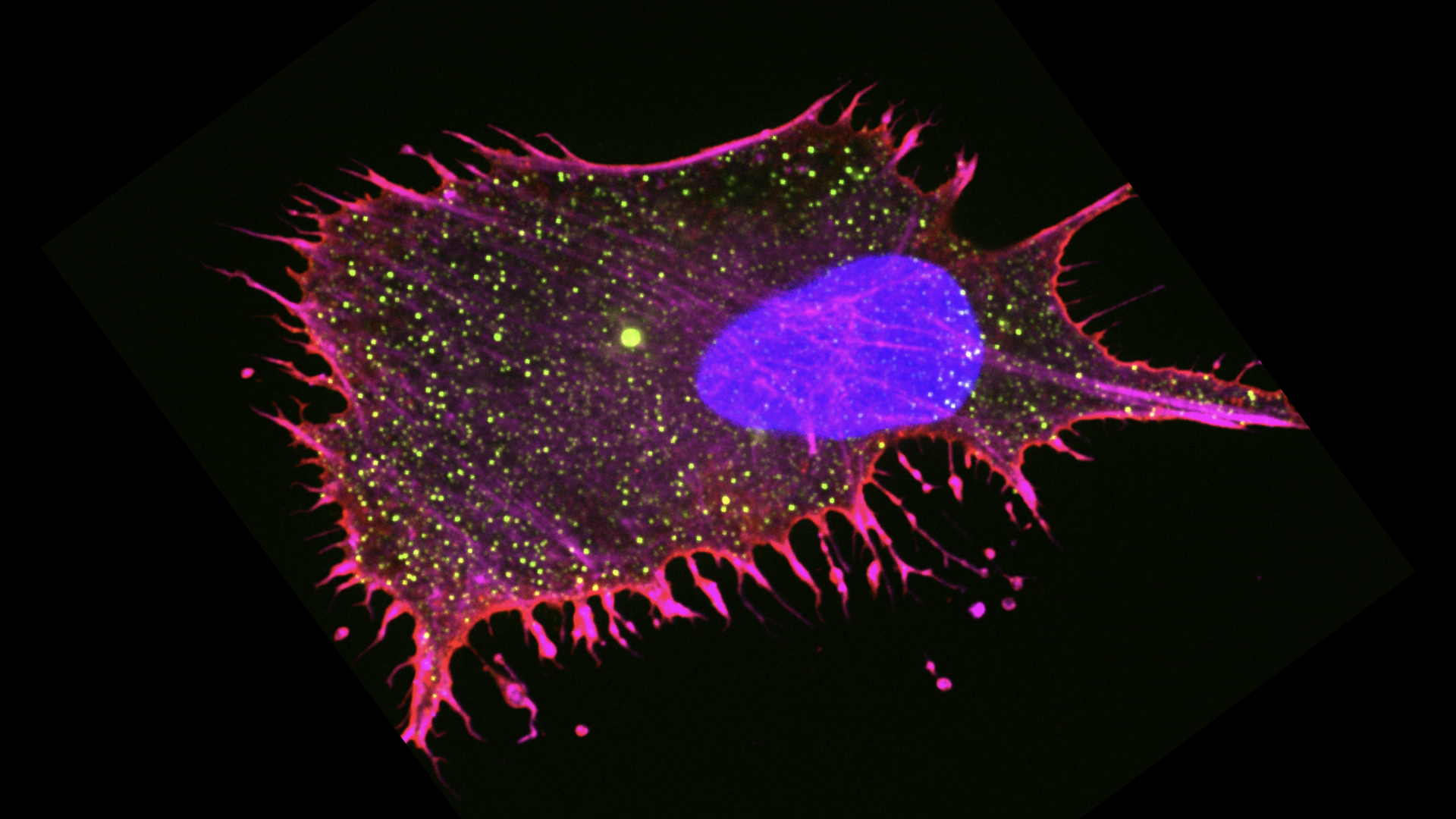
HOW DO CANCER CELLS ADAPT TO SURVIVE UNDER PRESSURE?
Physical pressure is crucial for cancer biology, but its effects remain poorly understood. When solid tumors grow confined within surrounding tissue, they build up pressure. Given that cells evolved to function in a stable mechanical environment, even slight changes in pressure perturb physiology. Normal cells and early stage cancer cells stop growing when pressure builds up, while advanced cancer cells are better able to grow and survive. This difference implies that cancer cells somehow adapt to physical pressure. This adaptation is of particular importance to pancreatic cancer, which builds up the most pressure of any tumor type. We are building devices to study how mechanical compression impacts the behavior and genome stability of cancer cells. We are leveraging machine learning to understand mitotic catastrophe under pressure and employing next-generation sequencing to understand genome rearrangements. We are screening for genes that enable cancer cells to survive under pressure and defining the mechanisms by which they cause mechanoadaptation in pancreatic cancer and Glioblastomas. We are also interested in identifying the early genetic events in the pathology of glioblastomas and their interplay with the stress.
The biophysics of neurodegeneration
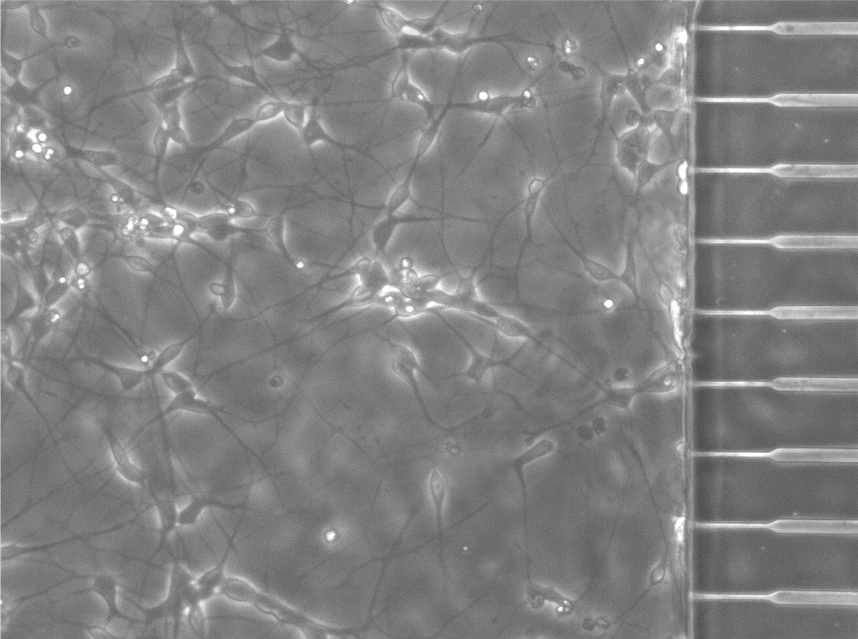
What initiates spontaneous neurodegeneration?
The aggregation and accumulation of disease-associated proteins such as TDP-43 is a notable neuropathological hallmark, yet we know little about why this highly abnormal event might occur. Rare genetic mutations implicate these proteins in disease, however more than 95% of neurodegeneration arises sporadically, and the mechanisms of sporadic disease remain unknown. We hypothesize that changes to the intracellular biophysical properties of neurons could be part of this mechanism.
Synthetic biology
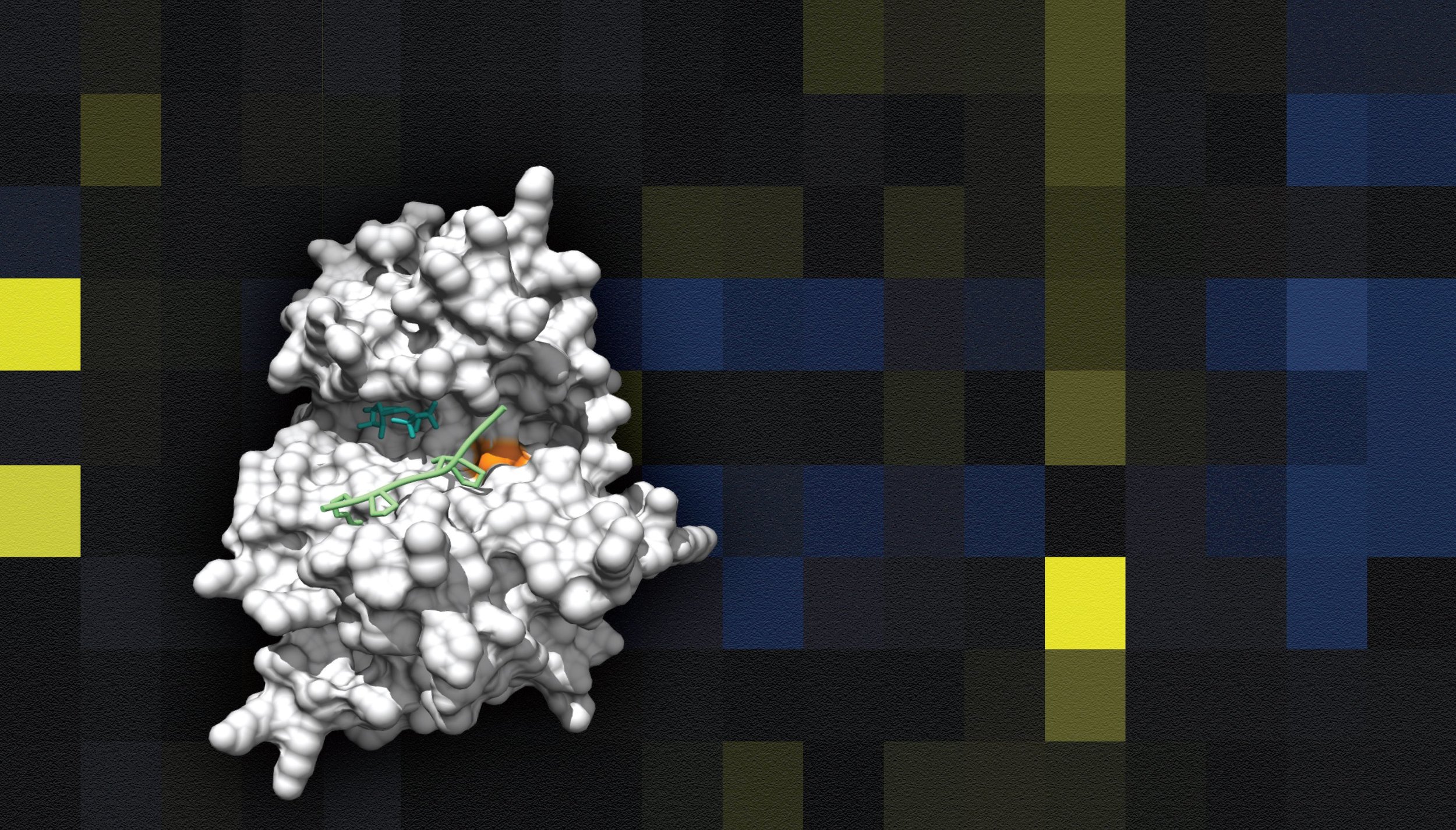
Can we understand biology by building systems from scratch?
Synthetic biology attempts to identify the building blocks of life. By attempting to engineer biological systems, we can begin to understand design rules and uncover new principles. If we are truly successful, we may even be able to create novel, useful biological functionality. These approaches are beginning to bear fruit with exciting new developments in genome editing and cellular engineering for medical, agricultural and industrial purposes. We are currently pursuing two synthetic biology projects:
syndrops - creating synthetic non-membrane bound compartments to study the effects of phase separation on signaling
In the highly crowded cellular environment, mRNAs and disordered proteins can condense together to form viscous liquid droplets. Biomolecules can be concentrated in or excluded from these compartments. Natural examples include stress granules, P-granules, nucleoli, and PML nuclear bodies. We are creating synthetic liquid droplets and targeting biochemical reactions within to try to understand some of the rules and consequences of this compartmentalization.
synHox - creating a synthetic hox locus to understand developmental transcriptional control
The Hox genes are a paradigm of spatio-temporal gene expression. If we can uyncover the rules that govern this precise regulation, we will have a far deeper understanding of epigenetic regulation. This knowledge will be extremely useful for stem-cell engineering and regenerative medicine. In collaboration with the Boeke and Mazzoni labs, we are synthesizing “big DNA” - we are making 100 kbp chunks of DNA and introducing them into stem cells. By "writing” an entire Hox locus from scratch and “rewriting” it however we want, we will uncover the rules of Hox expression.
Creating big-DNA from scratch (image by Sarah Richardson)
GP-write - a consortium to rewrite genomes.


